evolVienna Research Fields
1. Evolutionary Ecology
Evolutionary ecology studies the evolutionary consequences of complex selection pressures that arise in natural environments. The goal is to explain the evolution of the observed biodiversity. Research issues include adaptation to variable environments (e.g. phenotypic plasticity), interaction within species or among species (coevolution), and speciation theory. The evolVienna faculty address these questions on the organismal (Cremer, Gollmann, Hödl, Krenn, Paulus, Penn, Yamashita) as well as on the molecular level (Miller, Schleper, Stauffer), and with mathematical models (Dieckmann).
Principal Investigators: Sylvia Cremer, Ulf Dieckmann, Günter Gollmann, Walter Hödl, Harald W. Krenn, Wolfgang Miller, Hannes Paulus, Ovidiu Paun, Dustin Penn, Christa Schleper, Christian Stauffer, Kristin Tessmar, Nayuta Yamashita
2. Evolutionary Conservation Biology
Several research groups use an evolutionary perspective to address applied problems about conservation and restauration of biodiversity. The field comprises conservation genetics with a focus on the maintenance of genetic diversity (Burger, Haring) and ecological studies on the evolutionary response to human impact (fisheries induced evolution, Dieckmann).
Principal Investigators: Pamela Burger, Ulf Dieckmann, Elisabeth Haring, Elmira Mohandesan, Silvio Schueler

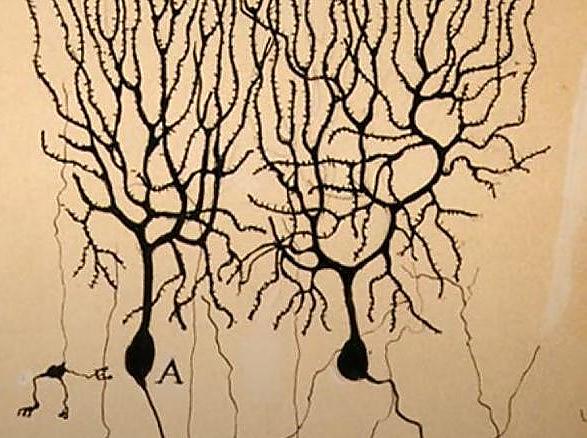
3. Evolution of the Nervous System, Behavior and Cognition
Research on the evolution of the nervous system, especially on cognitive abilities (such as speech) and behavioral patterns (including the evolution of cooperation) has a strong tradition in Vienna. Empirical groups study a large range of organisms from Drosophila (Miller), mites (Schausberger) and the bristle worm Platynereis (Raible, Tessmar) to vertebrates including fish, amphibians, birds, primates (Hödl, Fitch, Penn, Raible, Tessmar, Yamashita), and humans (Fitch). Modeling approaches rely on evolutionary game theory (Dieckmann, Hofbauer, Sigmund).
Principal Investigators: Ulf Dieckmann, Tecumseh Fitch, Walter Hödl, Josef Hofbauer, Wolfgang Miller, Dustin Penn, Florian Raible, Peter Schausberger, Karl Sigmund, Oleg Simakov, Kristin Tessmar, Nayuta Yamashita

4. Phylogenetics and Phylogeography
The reconstruction of the tree of life is one of the fundamental problems of evolutionary research. Phylogeography connects the problem of common decent with the patterns of contemporary geographical distribution of populations. Groups of evolVienna work on theoretical method development (von Haeseler) and applications across various taxonomic levels and geographic regions (Haring, Pass, Paulus, Schneeweiss, Stauffer, Vogl).
Principal Investigators: Arndt von Haeseler, Elisabeth Haring, Günther Pass, Hannes Paulus, Ovidiu Paun, Gerald Schneeweiss, Silvio Schueler, Oleg Simakov, Christian Stauffer, Claus Vogl

5. Experimental Evolution
Experimental evolution is the study of of evolutionary processes under controlled laboratory conditions under well-defined selective conditions. By exposing individuals to different selective environments (laboratory natural selection) or by directly selecting individuals for particular trait values (artificial selection) the experimenter can study the overall evolvability of traits, how focal traits and correlated traits respond to selection over time, and how genetic correlations constrain the evolutionary response. Moreover, by combining experimental evolution with genetic or genomic trait mapping methods the researcher can determine the genetic architecture underlying evolutionary change. The evolVienna faculty use this powerful experimental framework in microbes (Heidenreich, Schlötterer) and in Drosophila (Miller, Schlötterer) to address a range of questions related to adaptation and the functional significance of natural variation. Other groups (Cremer) study host-parasite coevolution in ant – bacteria/fungi systems.
Principal Investigators: Sylvia Cremer, Erich Heidenreich, Wolfgang Miller, Christian Schlötterer

6. Evo-Devo and Biometrics
Evolutionary developmental biology studies the evolution of developmental processes. Research issues include the evolutionary origin of body plans and the conditions for the evolution of modularity, organismal complexity, or novelty (evolutionary innovations). evolVienna scientists approach these questions on the phenotypic level (Müller, Metscher), where biometrics offers the statistical tools to quantify differences for morphological characters (Nemeschkal, Mitteröcker) or on the level of underlying genes (Eckhart, Raible, Technau, Tessmar).
Principal Investigators: Leopold Eckhart, Brian Metscher, Philipp Mitteröcker, Gerd B. Müller, Hans-Leo Nemeschkal, Florian Raible, Oleg Simakov, Ulrich Technau
7. Quantitative and Functional Genetics
The evolution of quantitative traits and the analysis of variance components is a classic area of evolutionary research. QTL (quantitative trait loci) and high resolution mapping methods aim to link phenotypic variation to the underlying genes. Complementing this approach, modern genetic tools facilitate direct functional tests of natural genetic variation observed among populations. Research in evolVienna addresses these issues using a range of approaches from theoretical modeling (Barton, Bürger) to empirical analysis linking phenotype to genotype (Nemeschkal), or focusing on particular candidate genes (Hauser, Nordborg, Raible, Schleper, Schlötterer, Technau).
Principal Investigators: Nick Barton, Reinhard Bürger, Marie-Theres Hauser, Hans-Leo Nemeschkal, Magnus Nordborg, Florian Raible, Christa Schleper, Christian Schlötterer, Silvio Schueler, Ulrich Technau, Kristin Tessmar
8. Population Genetics
The discipline that emerged from the modern synthesis of Darwinism with Mendelian inheritance studies the change of allele frequencies in populations under the evolutionary forces mutation, selection, recombination, drift, and gene-flow. Vienna hosts a large group of researchers in population genetics theory (Barton, Bürger, Futschik, Haeseler, Hermisson, Nordborg, Schuster, Vogl). Empirical population genetic research addresses various model and non-model organisms, including E. Coli, Drosophila (Haring, Schlötterer), Arabidopsis (Hauser, Nordborg), willows (Hauser), fire-bellied toads (Barton, Gollmann), camels (Burger), and humans.
Principal Investigators: Nick Barton, Pamela Burger, Reinhard Bürger, Andreas Futschik, Günter Gollmann, Arndt von Haeseler, Elisabeth Haring, Marie-Theres Hauser, Joachim Hermisson, Elmira Mohandesan, Magnus Nordborg, Christian Schlötterer, Silvio Schueler, Peter Schuster, Kristin Tessmar, Claus Vogl

9. Molecular Evolution and Genomics
Molecular evolutionary genetics uses comparative or polymorphism data to study evolution at the level of the DNA sequence. Studies may focus on candidate genes, but next generation sequencing technology allows also for genome-wide approaches (comparative and population genomics). Research questions range from the estimation of key population genetic parameters to the detection of the molecular loci of positive or negative selection in the history of a population. A large number of research groups in Vienna are involved in the production and analysis of data from various organisms from Bacteria and Archaea (Horn, Schleper) over budding yeast (Heidenreich) to the annelid worm Platynereis dumerilii (Raible), the sea anemone Nematostella vectensis (Technau), the bobtail squid Euprymna scolopes (Oleg Simakov), Drosophila (Miller, Schlötterer, Stauffer), Arabidopsis (Hauser, Nordborg), parasitic flowering plants (Schneeweiss) and higher primates (Eckhart). This empirical work is complemented by the development of stochastic models and statistical techniques needed for the interpretation of the data (Barton, Futschik, Haeseler, Hermisson, Nordborg, Vogl).
Principal Investigators: Nick Barton, Leopold Eckhart, Andreas Futschik, Elisabeth Haring, Marie-Theres Hauser, Arndt von Haeseler, Erich Heidenreich, Joachim Hermisson, Matthias Horn, Wolfgang Miller, Elmira Mohandesan, Magnus Nordborg, Ovidiu Paun, Florian Raible, Christa Schleper, Christian Schlötterer, Gerald Schneeweiss, Christian Stauffer, Oleg Simakov, Ulrich Technau, Kristin Tessmar, Claus Vogl

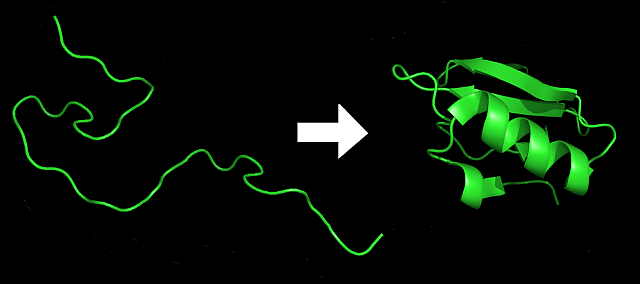
10. Sequence Evolution and Folding Landscapes
The secondary structure of RNA or the tertiary structure of amino acid sequences (proteins) constitutes a molecular phenotype that is directly determined by the sequence genotype. Selection on the structure then results in complex fitness landscapes for the evolution of the molecular sequence. Several evolVienna research groups study this relation for RNA, where folding algorithms based on biochemical principles and physical rules give rise to a computable sequence-to-structure map (Flamm, Hofacker, Schuster). The problem is even more complex for amino acid sequences, where primary, secondary, and tertiary structure contribute to complex patterns of constraints on the sequence level (Haeseler).
Principal Investigators: Christoph Flamm, Arndt von Haeseler, Ivo Hofacker, Peter Schuster
11. Paleogenomics
Ancient DNA (aDNA) is a new field of science that began in the early 1980s, based on extraction and analysis of small fragments of mitochondrial DNA (mtDNA) from quagga, an extinct subspecies of the plains zebra. Ancient DNA can be recovered and analyzed from museum specimens, fossil remains, archaeological and paleontological finds. Over the past decades, aDNA research has entered the new era of genomics - "Paleogenomics" - due to recent advances in laboratory methods, next-generation DNA sequencing technology and bioinformatic tools. At present, whole genomes from ancient anatomically modern humans, archaic hominins, ancient pathogens and various non-human animals have been successfully sequenced. Today' application of ancient DNA as a powerful research tool is covering a broad range of evolutionary questions in the context of dynamic processes of migration, adaptation, admixture, domestication, and potential causes of extinctions in past species. Research groups in Vienna (Pinhasi, Mohandesan, Haring, Burger, Wallner) are focusing on various evolutionary topics on humans and animals, using ancient DNA.
Principal Investigators: Pamela Burger, Elisabeth Haring, Elmira Mohandesan, Ron Pinhasi, Barbara Wallner
